Note
Go to the end to download the full example code.
Fetch Your First Study¶
Download a sample dataset, load it as an events DataFrame, and explore the event structure — the same workflow that works for all neuralfetch studies.
Discover available studies¶
ns.Study.catalog() lists every study registered in neuralfetch,
including full datasets and their lightweight sample variants.
import collections
import neuralset as ns
all_studies = ns.Study.catalog()
print(f"{len(all_studies)} studies available (full + sample variants)")
modalities = collections.Counter(
nt for cls in all_studies.values() for nt in cls.neuro_types()
)
print("By modality:", dict(modalities))
132 studies available (full + sample variants)
By modality: {'Eeg': 115, 'Fmri': 6, 'Meg': 8, 'Ieeg': 1, 'Fnirs': 1}
Inspect a study’s metadata¶
Every study exposes class-level metadata — no download needed.
Study = all_studies["Grootswagers2022Human"]
print(f"Description: {Study.description[:100]}...")
print(f"Neuro types: {Study.neuro_types()}")
info = Study._info
print(f"Subjects: {info.num_subjects}, timelines: {info.num_timelines}")
Description: EEG recordings from 50 participants watching still images....
Neuro types: frozenset({'Eeg'})
Subjects: 50, timelines: 50
Load a sample and preview events¶
Sample studies are small subsets that download automatically. They use the same API as full datasets — only the study name differs.
Here we use Fake2025Meg which requires no download and
demonstrates the same events structure you’ll see with real data.
0%| | 0/2 [00:00<?, ?it/s]
50%|█████ | 1/2 [00:00<00:00, 3.18it/s]
100%|██████████| 2/2 [00:00<00:00, 3.18it/s]
100%|██████████| 2/2 [00:00<00:00, 3.18it/s]
Loaded 1588 events from 2 subject(s)
type start duration subject
0 Meg 42.955971 277.715346 Fake2025Meg/sample0
1 Text 46.578924 210.637433 Fake2025Meg/sample0
2 Audio 46.578924 210.637375 Fake2025Meg/sample0
3 Sentence 46.578924 2.264551 Fake2025Meg/sample0
4 Word 46.578924 0.200000 Fake2025Meg/sample0
5 Stimulus 46.578924 0.006660 Fake2025Meg/sample0
6 Word 47.191629 0.200000 Fake2025Meg/sample0
7 Image 47.191629 0.200000 Fake2025Meg/sample0
8 Stimulus 47.191629 0.006660 Fake2025Meg/sample0
9 Word 47.897572 0.200000 Fake2025Meg/sample0
10 Stimulus 47.897572 0.006660 Fake2025Meg/sample0
11 Word 48.643475 0.200000 Fake2025Meg/sample0
Explore event types¶
Every study returns the same schema: a flat DataFrame where brain
recordings, stimuli, text, and annotations share the same rows,
distinguished by the type column.
print("Event types:")
print(events["type"].value_counts().to_string())
Event types:
type
Word 576
Stimulus 576
Image 286
Sentence 144
Meg 2
Text 2
Audio 2
Each type has its own extra columns:
Meg: ['filepath']
Text: ['text']
Audio: ['filepath']
Sentence: ['text']
Word: ['text']
Stimulus: ['code']
Image: ['filepath']
Visualise the timeline¶
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(10, 3))
types = events["type"].unique()
colors = plt.cm.Set2(range(len(types)))
for i, t in enumerate(types):
sub = events[events["type"] == t]
ax.barh(
i,
sub["duration"].fillna(0.1),
left=sub["start"],
height=0.6,
label=t,
color=colors[i],
)
ax.set_yticks(range(len(types)))
ax.set_yticklabels(types)
ax.set_xlabel("Time (s)")
ax.set_title("Fake2025Meg — event timeline (subject 0)")
ax.legend(loc="upper right")
plt.tight_layout()
plt.show()
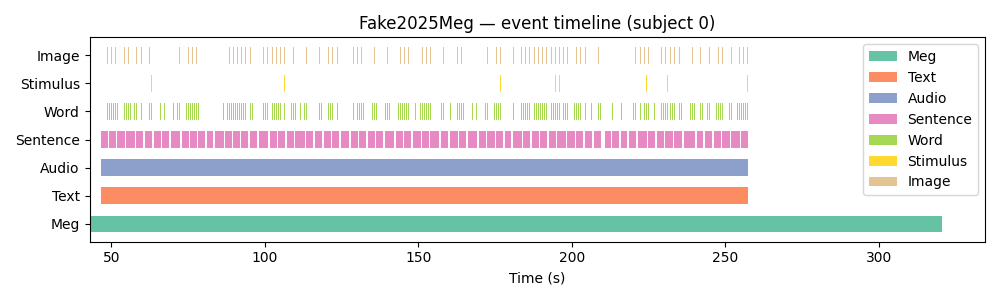
Using a real sample dataset¶
Replace Fake2025Meg with any sample name from the catalog to run
on real data. All sample studies download their files automatically:
See Sample Datasets for all available sample datasets and ready-to-paste loading snippets.
Next: Anatomy of a Study — timelines, event anatomy, and composing studies with transforms.
Total running time of the script: (0 minutes 1.714 seconds)